HEASOFT, CALDB and NuSTAR data reduction#
contact: menglei.zhou@astro.uni-tuebingen.de
0. Prerequisites#
0.1. Installation of HEAsoft#
excute
wget https://heasarc.gsfc.nasa.gov/FTP/software/lheasoft/lheasoft6.36/heasoft-6.36src.tar.gz in your preferred installation directory
and follow the installation guidance at https://heasarc.gsfc.nasa.gov/docs/software/lheasoft/linux.html. It shall take a few hours to finish.
We strongly recommend you to compile the source code to proceed (in order to use local models)!
0.2. Installing the calibration database ($CALDB)#
follow the guidance at https://heasarc.gsfc.nasa.gov/docs/heasarc/caldb/caldb_install.html to install the calibration database software.
cd $CALDBfind the calibration files of NuSTAR mission at https://heasarc.gsfc.nasa.gov/docs/heasarc/caldb/caldb_supported_missions.html
run
wget https://heasarc.gsfc.nasa.gov/FTP/caldb/data/nustar/fpm/goodfiles_nustar_fpm_20251027.tar.gzto download the NuSTAR calibration files.tar -zxf goodfiles_nustar_fpm_20251027.tar.gzfor future practice, you may also download the calibration files:
NICER:
wget https://heasarc.gsfc.nasa.gov/FTP/caldb/data/nicer/xti/goodfiles_nicer_xti_20240206.tar.gz;IXPE:
wget https://heasarc.gsfc.nasa.gov/FTP/caldb/data/ixpe/gpd/goodfiles_ixpe_gpd_20250225.tar.gz&wget https://heasarc.gsfc.nasa.gov/FTP/caldb/data/ixpe/xrt/goodfiles_ixpe_xrt_20241028.tar.gzand unzip them in $CALDB. The calibration files will be delivered to the proper position automatically.
0.3. Installing DS9 (for image handling)#
go to https://sites.google.com/cfa.harvard.edu/saoimageds9 and follow the installation guidance
1. Accessing Observational Data from High-Energy Astrophysics Archives#
go to https://heasarc.gsfc.nasa.gov/cgi-bin/W3Browse/w3browse.pl and search for your interested source
e.g., the famous galactic black hole binary: Cygnus X-1
1.1. Download the level 1 data (the preprocessed data of your observation)#
wget -q -nH --no-check-certificate --cut-dirs=6 -r -w1 -l0 -c -N -np -R 'index*' -erobots=off --retr-symlinks https://heasarc.gsfc.nasa.gov/FTP/nustar/data/obs/10/9//91002320002/
2. Run the pipeline to produce the NuSTAR level 2 data (clean photon event lists)#
nupipeline indir=/data/to/your/path/91002320002 outdir=/data/to/your/path/91002320002/out steminputs=nu91002320002
This step is quite time-consuming and may take several hours to complete. But don’t worry — we’ve already prepared the processed results for you! You can download them here: https://drive.google.com/file/d/1MqZozxRo7Rp7XxY6zoU5GqNI5Fz_Volm/view?usp=sharing
2.1. The FITS file structure (HDU := Header + Data)#
2.2. Header#
2.3. Data#
3. Using DS9 to extract the source region and the background region#
ds9 out/nu91002320002A01_cl.evt and ds9 out/nu91002320002B01_cl.evt
Please refer to https://heasarc.gsfc.nasa.gov/docs/nustar/analysis/nustar_quickstart_guide.pdf as a quick-start guide and https://heasarc.gsfc.nasa.gov/docs/nustar/analysis/nustar_swguide.pdf for further details.
In brief, the source region should encompass most of the emission area when viewed on a logarithmic color scale, while the background region should be as large as possible—excluding any part of the source—and selected from the same detector quadrant.
4. Extract the spectra and light curves (level 3 data that can be used directly for scientific purposes.)#
nuproducts srcregionfile=./out/src.reg bkgregionfile=./out/bkg.reg indir=./out outdir=./products instrument=FPMA steminputs=nu91002320002 bkgextract=yes
nuproducts srcregionfile=./out/src.reg bkgregionfile=./out/bkg.reg indir=./out outdir=./products instrument=FPMB steminputs=nu91002320002 bkgextract=yes
the whole process can take 10+ minutes to complete. Or you can get the products via https://drive.google.com/file/d/1vXcH7zNuNQT6ojTcW6iQM6l2KcuxV5FZ/view?usp=sharing
5. Check the structure of your final scientific products#
We can use fv to quickly check the structure of FITS files.
fv NAME_OF_YOUR_INTERESTED_FILE.fits (.evt .lc .pha .rmf .arf ...)
5.1. The structure of light curve files#
the correlation between rate_orginal and rate:#
5.2. The structure of the spectrum#
Next, we’ll handle more complex FITS files—most of these operations can be done conveniently with astropy.
import numpy as np
import astropy.io.fits as fits
import matplotlib.pyplot as plt
5.2.1. Spectrum file#
spec = fits.open('../data/nu91002320002A01_sr.pha')
spec.info()
Filename: ../data/nu91002320002A01_sr.pha
No. Name Ver Type Cards Dimensions Format
0 PRIMARY 1 PrimaryHDU 504 (105, 105) float32
1 SPECTRUM 1 BinTableHDU 468 4096R x 2C [J, J]
2 GTI 1 BinTableHDU 57 13R x 2C [1D, 1D]
3 REG00101 1 BinTableHDU 84 1R x 6C [1PD(1), 1PD(1), 16A, 1PD(1), 1PD(0), 1PI(1)]
bkg_spec = fits.open('../data/nu91002320002A01_bk.pha')
bkg_spec.info()
Filename: ../data/nu91002320002A01_bk.pha
No. Name Ver Type Cards Dimensions Format
0 PRIMARY 1 PrimaryHDU 504 (143, 143) float32
1 SPECTRUM 1 BinTableHDU 468 4096R x 2C [J, J]
2 GTI 1 BinTableHDU 57 13R x 2C [1D, 1D]
3 REG00101 1 BinTableHDU 84 1R x 6C [1PD(1), 1PD(1), 16A, 1PD(1), 1PD(0), 1PI(1)]
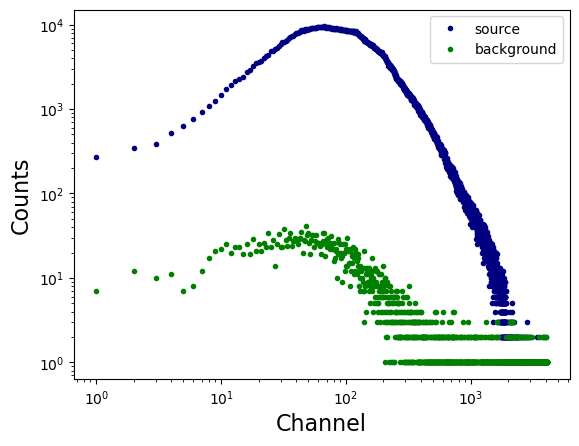
x = spec['SPECTRUM'].data['CHANNEL']
y = spec['SPECTRUM'].data['COUNTS']
plt.loglog(x, y, linestyle = 'none', marker = '.', markerfacecolor = 'navy', markeredgecolor = 'navy', label = 'source')
x = bkg_spec['SPECTRUM'].data['CHANNEL']
y = bkg_spec['SPECTRUM'].data['COUNTS']
plt.loglog(x, y, linestyle = 'none', marker = '.', markerfacecolor = 'green', markeredgecolor = 'green', label = 'background')
plt.xlabel('Channel', fontsize = 16)
plt.ylabel('Counts', fontsize = 16)
plt.legend()
plt.show()
5.2.2. Response Matrix Files (RMFs)#
resp = fits.open('../data/nu91002320002A01_sr.rmf')
resp.info()
Filename: ../data/nu91002320002A01_sr.rmf
No. Name Ver Type Cards Dimensions Format
0 PRIMARY 1 PrimaryHDU 14 ()
1 EBOUNDS 1 BinTableHDU 58 4096R x 3C [J, E, E]
2 MATRIX 1 BinTableHDU 52 4096R x 6C [E, E, I, PI(117), PI(117), PE(4027)]
plt.plot(resp['EBOUNDS'].data['CHANNEL'], resp['EBOUNDS'].data['E_MIN'])
plt.show()
print(resp['EBOUNDS'].data['E_MAX'] - resp['EBOUNDS'].data['E_MIN'])
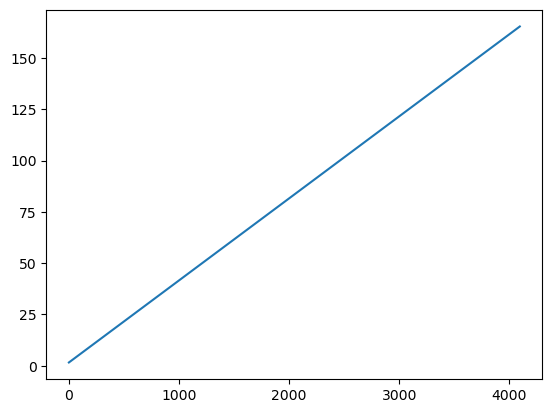
[0.03999996 0.03999996 0.04000008 ... 0.03999329 0.03999329 0.04000854]
x = resp['EBOUNDS'].data['CHANNEL']
idx_1 = 50
idx_2 = 500
idx_3 = 1000
idx_4 = 1500
idx_5 = 2000
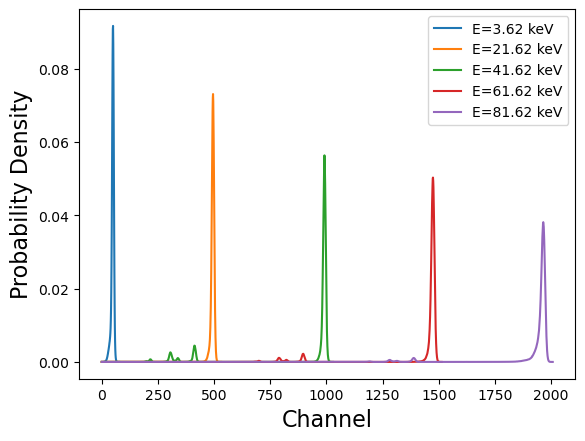
y = resp['MATRIX'].data['MATRIX'][idx_1]/np.sum(resp['MATRIX'].data['MATRIX'][idx_1])
plt.plot(y, label = 'E={0:0.2f} keV'.format(resp['EBOUNDS'].data['E_MIN'][idx_1] + 0.02))
y = resp['MATRIX'].data['MATRIX'][idx_2]/np.sum(resp['MATRIX'].data['MATRIX'][idx_2])
plt.plot(y, label = 'E={0:0.2f} keV'.format(resp['EBOUNDS'].data['E_MIN'][idx_2] + 0.02))
y = resp['MATRIX'].data['MATRIX'][idx_3]/np.sum(resp['MATRIX'].data['MATRIX'][idx_3])
plt.plot(y, label = 'E={0:0.2f} keV'.format(resp['EBOUNDS'].data['E_MIN'][idx_3] + 0.02))
y = resp['MATRIX'].data['MATRIX'][idx_4]/np.sum(resp['MATRIX'].data['MATRIX'][idx_4])
plt.plot(y, label = 'E={0:0.2f} keV'.format(resp['EBOUNDS'].data['E_MIN'][idx_4] + 0.02))
y = resp['MATRIX'].data['MATRIX'][idx_5]/np.sum(resp['MATRIX'].data['MATRIX'][idx_5])
plt.plot(y, label = 'E={0:0.2f} keV'.format(resp['EBOUNDS'].data['E_MIN'][idx_5] + 0.02))
plt.legend()
# plt.yscale('log')
plt.xlabel('Channel', fontsize = 16)
plt.ylabel('Probability Density', fontsize =16)
plt.show()
5.2.3. Ancillary Response Files (ARFs)#
arf = fits.open('../data/nu91002320002A01_sr.arf')
arf.info()
Filename: ../data/nu91002320002A01_sr.arf
No. Name Ver Type Cards Dimensions Format
0 PRIMARY 1 PrimaryHDU 12 ()
1 SPECRESP 1 BinTableHDU 88 4096R x 3C [E, E, E]
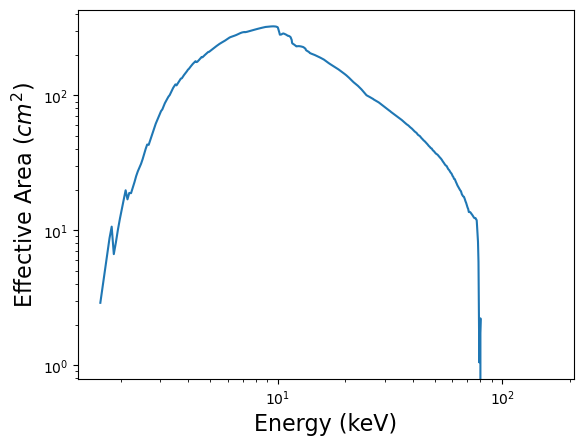
x = 0.5*(arf['SPECRESP'].data['ENERG_LO'] + arf['SPECRESP'].data['ENERG_HI'])
y = arf['SPECRESP'].data['SPECRESP']
plt.loglog(x, y)
plt.xlabel('Energy (keV)', fontsize = 16)
plt.ylabel('Effective Area ($cm^2$)', fontsize = 16)
plt.show()
5.3. Group your spectra so that Gaussian statistics can be validly applied#
ftgrouppha nu91002320002A01_sr.pha nu91002320002A01_sr_grp.pha grouptype=optmin groupscale=30 respfile=nu91002320002A01_sr.rmf clobber=yes
ftgrouppha nu91002320002B01_sr.pha nu91002320002B01_sr_grp.pha grouptype=optmin groupscale=30 respfile=nu91002320002B01_sr.rmf clobber=yes
please read https://heasarc.gsfc.nasa.gov/docs/software/lheasoft/help/ftgrouppha.html for further details.
6. Fit your spectra with XSPEC#
Simply enter xspec in the command prompt to start XSPEC.
We often adopt the elemental abundances of the Galaxy as defined by the solar abundances measured by Wilms et al. (2000). To set this in XSPEC, type abund wilm in the XSPEC prompt.
Type query yes to prevent the program from requesting further confirmations until you obtain the best fit.
6.1. Loading the data into XSPEC#
data 1:1 nu91002320002A01_sr_grp.pha
data 2:2 nu91002320002B01_sr_grp.pha
(The number before the colon specifies how many model sets you want to fit, while the number after the colon indicates the spectrum number. Try running data 1:1 nu91002320002A01_sr_grp.pha and data 1:2 nu91002320002B01_sr_grp.pha,
then define a model to observe the difference.)
ignore 1-2: **-3.0 79.0-** (This command excludes the poorly calibrated energy ranges. Note: Always include the decimal point — otherwise, XSPEC will interpret the value as a channel number instead of an energy value.)
6.2. Fitting#
The model is calculated in flux space and then transformed into channel space using the response matrix and the effective area.
C(I): the expectation of the observed count spectrum;
S(E): the (defined) intrinsic source spectrum (in flux space);
R(I, E): the normalized instrumental response matrix;
A(E): the effective area of the instrument;
B(I): the background spectrum.
Type model constant*TBabs*powerlaw in XSPEC.
constant — accounts for the flux normalization between the two NuSTAR instruments, FPMA and FPMB.
TBabs — represents Galactic absorption.
powerlaw — describes the spectral continuum, which is primarily dominated by inverse Compton scattering.
Simply type fit once all parameters have been set.
cpd /xs or cpd /xw to set the proper plotting window
plot uf del: this is the command to show the unfolded spectrum (in flux space) and the \(\chi\) residuals.
We can see a very porminent iron line and some humps at 20-30 keV.
Or we can try model constant*TBabs*(powerlaw + diskbb) to consider the contribution from the accretion disk.
The iron line is still there, not resolved by the thermal disk model diskbb.
Or we can try model constant*TBabs*(powerlaw + diskbb + gaussian) to cover the Gaussian line at 6-7 keV.
What should be the best model? We can try with some physical models…
constant*TBabs*(nthcomp + relxill + gaussian), how about this?
The details of the relxill model family will be discussed in the following lectures!
